BREAKING THE ANTIBIOTIC RESISTANCE: EZGİ KARACA
Transposase-DNA Complex Structures Reveal Mechanisms for Conjugative Transposition of Antibiotic Resistance
EMBL (Barabas Lab) and IBG (Karaca Lab) researchers have recently revealed the structural basis of a prominent antibiotic resistance transfer mechanism. Targeting this mechanism entails the potential to fight against a major global health threat, antibiotic resistant bacteria.
This collaborative work presents the first complex structure, unraveling how resistance carrying transposons are inserted into DNA. Dissecting this structure paves the way for the development of system-specific drugs, which can directly block the resistance carrying transposon transfer mechanism. This work is published in Cell.

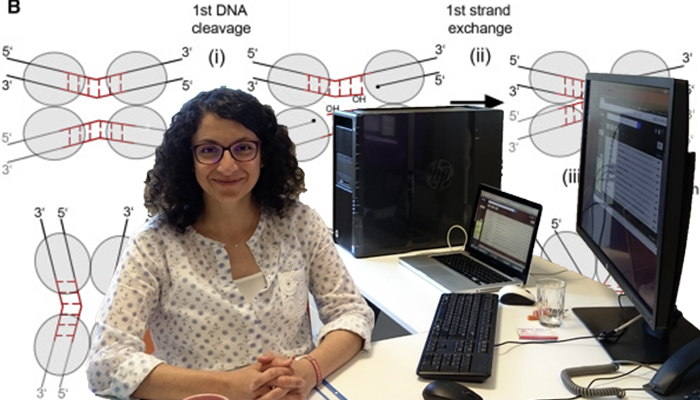
The proposed mechanism for the resistance carrying transposon insertion (adapted from Figure 1 of the Rubio-Cosials et al.): (i) two proteins cleave one strand of each dsDNA, (ii) the 5′ ends swap places and resolve the protein-DNA link on the partner strands, generating a four-way Holliday junction (HJ) intermediate; (iii) isomerization activates the second protein pair to cleavage (iv) and exchange (v) the other DNA strands, generating the final recombined products.
Anna Rubio-Cosials, Eike C. Schulz, Lotte Lambertsen, Georgy Smyshlyaev, Carlos Rojas-Cordova, Kristoffer Forslund, Ezgi Karaca, Aleksandra Bebel, Peer Bork, Orsolya Barabas, Transposase – DNA complex structures reveal mechanisms for conjugative transposition of antibiotic resistance. Cell, published online 15 March, 2018. DOI: 10.1016/j.cell.2018.02.032
EMBL press release: https://news.embl.de/science/potential-new-way-limit-antibiotic-resistance-spreading/