Kucuk Lab. on Lymphoid Cancer Genomics
RESEARCH INTERESTS
Our current research mainly focuses on the identification and characterization of novel mutations and epigenetic alterations of tumor suppressor genes or oncogenes in lymphoid cancers. Our interest mainly resides in lymphomas (e.g. follicular lymphoma) and other lymphoid cancers (e.g. multiple myeloma) which are neoplastic diseases having a high degree of genetic heterogeneity.
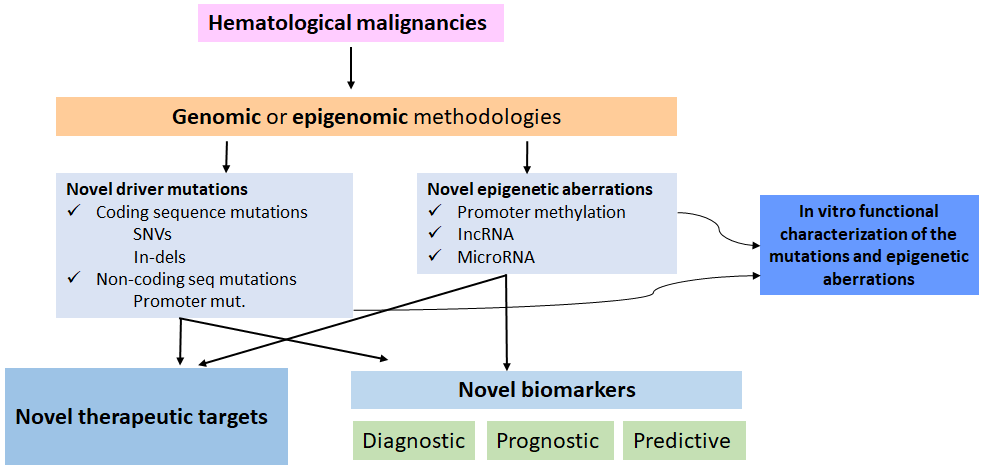
We frequently use next generation sequencing (NGS) based technologies (e.g. WES, RNA-Seq, targeted NGS) to generate a candidate list of aberrant genes using panels of lymphoma patient samples. We then cross-validate mutations that are identified through NGS methodologies by applying Sanger sequencing. Moreover, our studies focus on genome-wide or loci-specific methylation changes in cancer samples. Altogether, we characterize these aberrations using a combination of genetic, epigenetic and molecular biology assays. In general, our studies follow a wholistic approach which involves the cooperation of a variety interrelated biomedical disciplines such as pathology, functional genomics, and computational bioinformatics. Our eventual goal is to identify and characterize genetic and epigenetic abnormalities in individual cancer patients to translate the discoveries as personalized medicine.
Group Members

Kucuk Lab. on Lymphoid Cancer Genomics
Former Members

Can KÜÇÜK
Research Group Leader
can.kucuk@ibg.edu.tr
+90 232 299 41
00
(5011)
0 232 299 41 51

Ayla ANAR
Research Technician
ayla.anar@ibg.edu.tr

Esra ESMERAY
PhD Student
esra.esmeray@ibg.edu.tr

Deniz KURŞUN
MSc Student
deniz.kursun@msfr.ibg.edu.tr

Burcu AKMAN
PhD Student
burcu.akman@ibg.edu.tr
0 (232) 299 _ _

Tevfik HATİPOĞLU
PhD Student
tevfik.hatipoglu@ibg.edu.tr

Xiaozhou HU
Visiting Researcher
xiaozhou.hu@ibg.edu.tr
+90 232 299 41
00
(4101)

Işıl KILIÇGÜN
MSc Student
isil.kilicgun@ibg.edu.tr

EZGİ BOYVATLI
MSc Student
ezgi.boyvatli@ibg.edu.tr

Arda CEYLAN
MSc Student
arda.ceylan@ibg.edu.tr

Ayla ANAR
Researcher
ayla.anar@ibg.edu.tr

Tevfik HATİPOĞLU
Post-Doc Researcher
tevfik.hatipoglu@ibg.edu.tr
News
A new international award from the American Society of Hematology to Assoc. Prof. Can Küçük
Assoc. Prof. Dr. Can Küçük earned 40.000 USD project award of the ASH Awards Supplement Program from the American Society of Hematology (ASH), which is the largest professional society of hematology in the world. The awardees for the ASH Supplement Award were selected among previous winners with ongoing projects of ASH awards in 5 different categories including prestigous awards such as the ASH Global Research Award and ASH Scholar Award. ASH Supplement Award is the 2nd award from the ASH in its 64 years of history to a scientist working in Turkey after the ASH Global Research Award that was also provided to Dr. Can Küçük in 2018. You can obtain more information regarding this award program through the following link: https://www.hematology.org/awards/career-enhancement-and-training/ash-awards-supplement-program
An international collaborative study from Lymphoid Cancer Genomics Research Group is highlighted in the Journal of Leukocyte Biology
IL2 signaling plays a crucial role during NK cell activation including proliferation as a response to virus-infected cells or neoplastic cells. Appropriate termination of IL2 signaling is required to control NK cell expansion and prevent NK cell malignancies. The molecular mechanisms responsible for termination of IL2 signaling were largely unknown. An international collaborative study from Dr. Küçük’s research group revealed that PRDM1 decreases sensitivity of NK cells to IL2 cytokine through direct transcriptional inhibition of IL2RA, thereby preventing uncontrolled NK cell expansion.
From Lymphoid Cancer Genomics Research Group Dr. Burcu Akman, Dr. Xiaozhou Hu, Tevfik Hatipoğlu, and Dr. Can Küçük contributed to this study, which has been highlighted in the recent issue of the Journal of Leukocyte Biology. For more information, please see the following link: https://jlb.onlinelibrary.wiley.com/
Selected Publications
Altun Z, Yuan H, Baran B, Aktaş S, Esmeray Sönmez E, Küçük C, Olgun N. Whole-exome sequencing reveals genetic variants in low-risk and high-risk neuroblastoma.. Gene. 2023 April ; 860 : 147233. doi:10.1016/j.gene.2023.147233.
Ahmet Şeyhanlı, Tevfik Hatipoğlu, Burcu Akman, Xiaozhou Hu, Güner Hayri Özsan, Can Küçük. Genetic and Epigenetic Analyses Reveal Transcriptional Silencing of SOX7 due to Deletion in a Multiple Myeloma Case with Double Relapse. International Journal of Hematology and Oncology. 2023 April ; 33 (1306-133X) : 054-056. doi:10.4999/uhod.236587.
Hatipoğlu T, Esmeray Sönmez E, Hu X, Yuan H, Danyeli AE, Şeyhanlı A, Önal-Süzek T, Zhang W, Akman B, Olgun A, Özkal S, Alacacıoğlu İ, Özcan MA, You H, Küçük C. Plasma Concentrations and Cancer-Associated Mutations in Cell-Free Circulating DNA of Treatment-Naive Follicular Lymphoma for Improved Non-Invasive Diagnosis and Prognosis.. Frontiers in Oncology. 2022 June ; 12 : 870487. doi:10.3389/fonc.2022.870487.
Esmeray Sönmez, Esra, Tevfik Hatipoğlu, Deniz Kurşun, Xiaozhou Hu, Burcu Akman, Hongling Yuan, Ayça Erşen Danyeli, İnci Alacacıoğlu, Sermin Özkal, Aybüke Olgun, Taner Kemal Erdağ, Hua You, and Can Küçük. Whole Transcriptome Sequencing Reveals Cancer-Related, Prognostically Significant Transcripts and Tumor-Infiltrating Immunocytes in Mantle Cell Lymphoma. Cells. 2022 October ; 11 (21) : 3394. doi:10.3390/cells11213394.
Mengyuan Dai, Miao Liu, Hua Yang, Can Küçük, Hua You. New insights into epigenetic regulation of resistance to PD-1/PD-L1 blockade cancer immunotherapy: mechanisms and therapeutic opportunities. Experimental hematology & oncology. 2022 November . doi:10.1186/s40164-022-00356-0.
Wang Y, Ba HJ, Wen XZ, Zhou M, Küçük C, Tamagnone L, Wei L, You H. A prognostic model for melanoma patients on the basis of immune-related lncRNAs.. Aging. 2021 March ; 13 . doi:10.18632/aging.202730.
Yılmaz H, Toy HI, Marquardt S, Karakülah G, Küçük C, Kontou PI, Logotheti S, Pavlopoulou A. In Silico Methods for the Identification of Diagnostic and Favorable Prognostic Markers in Acute Myeloid Leukemia.. International journal of molecular sciences. 2021 September ; 22 (17) : 9601. doi:10.3390/ijms22179601.
Küçük C. Genetic susceptibility to natural killer T-cell lymphoma.. The Lancet. Oncology. 2020 February ; 21 (2) : 196-197. doi:10.1016/S1470-2045(19)30820-4.
Esmeray E, Küçük C. Genetic alterations in B cell lymphoma subtypes as potential biomarkers for noninvasive diagnosis, prognosis, therapy, and disease monitoring.. Turkish journal of biology = Turk biyoloji dergisi. 2020 February ; 44 (1) : 1-14. doi:10.3906/biy-1908-23.
Akman B, Hu X, Liu X, Hatipoğlu T, You H, Chan WC, Küçük C. PRDM1 decreases sensitivity of human NK cells to IL2-induced cell expansion by directly repressing CD25 (IL2RA).. Journal of leukocyte biology. 2020 November . doi:10.1002/JLB.2A0520-321RR.
Küçük C, Wei L, You H. Indolent T-Cell Lymphoproliferative Disease of the GI Tract: Insights for Better Diagnosis, Prognosis, and Appropriate Therapy.. Frontiers in oncology. 2020 August ; 10 : 1276. doi:10.3389/fonc.2020.01276.
Küçük C, Wang J, Xiang Y, You H. Epigenetic aberrations in natural killer/T-cell lymphoma: diagnostic, prognostic and therapeutic implications.. Therapeutic advances in medical oncology. 2020 February ; 12 : 1758835919900856. doi:10.1177/1758835919900856.
Kursun, Deniz; Kucuk, Can. Systematic analysis of the frequently amplified 2p15-p16.1 locus reveals PAPOLG as a potential proto-oncogene in follicular and transformed follicular lymphoma. TURKISH JOURNAL OF BIOLOGY. 2019 ; 43 (2) . doi:10.3906/biy-1810-2.
Wan Y, He D, Ye Y, Zhang W, Zhao S, Long Y, Chen M, Küçük C. Primary cardiac diffuse large B-cell lymphoma with concurrent high MYC and BCL2 expression in an immunocompetent Chinese elderly woman. Cardiovasc Pathol. 2017 November ; 31 : 54-56. doi:10.1016/j.carpath.2017.07.006.
Hu X, Baytak E, Li J, Akman B, Okay K, Hu G, Scuto A, Zhang W, Küçük C. The relationship of REL proto-oncogene to pathobiology and chemoresistance in follicular and transformed follicular lymphoma. Leuk Res. 2017 March ; 54 : 30-38. doi:10.1016/j.leukres.2017.01.001.
Baytak E, Gong Q, Akman B, Yuan H, Chan WC, Küçük C. Whole transcriptome analysis reveals dysregulated oncogenic lncRNAs in natural killer/T-cell lymphoma and establishes MIR155HG as a target of PRDM1.. Tumour biology : the journal of the International Society for Oncodevelopmental Biology and Medicine. 2017 May ; 39 (5) : 1010428317701648. doi:10.1177/1010428317701648.
Gao, Li-Min; Zhao, Sha; Liu, Wei-Ping; Zhang, Wen-Yan; Li, Gan-Di; Kucuk, Can; Hu, Xiao-Zhou; Chan, Wing C.; Tang, Yuan; Ding, Wen-Shuang; Yan, Jia-Qi; Yao, Wen-Qing; Wang, Jian Chao. Clinicopathologic Characterization of Aggressive Natural Killer Cell Leukemia Involving Different Tissue Sites. AMERICAN JOURNAL OF SURGICAL PATHOLOGY. 2016 June ; 40 (6) : 836-846. doi:10.1097/PAS.0000000000000634.
Kucuk, Can; Hu, Xiaozhou; Gong, Qiang; Jiang, Bei; Cornish, Adam; Gaulard, Philippe; McKeithan, Timothy; Chan, Wing C.. Diagnostic and Biological Significance of KIR Expression Profile Determined by RNA-Seq in Natural Killer/T-Cell Lymphoma. AMERICAN JOURNAL OF PATHOLOGY. 2016 June ; 186 (6) : 1435-1441. doi:10.1016/j.ajpath.2016.02.011.
Hu X, Chan WC, Kücük C. Generation of a genetically engineered aggressive nk-cell leukemia cell line with stable IL2 expression. Acta Medica International. 2015 July ; 2 (2) : 78-84. doi:10.5530/ami.2015.3.6.
Küçük C, Jiang B, Hu X, Zhang W, Chan JK, Xiao W, Lack N, Alkan C, Williams JC, Avery KN, Kavak P, Scuto A, Sen E, Gaulard P, Staudt L, Iqbal J, Zhang W, Cornish A, Gong Q, Yang Q, Sun H, d'Amore F, Leppä S, Liu W, Fu K, de Leval L, McKeithan T, Chan WC. Activating mutations of STAT5B and STAT3 in lymphomas derived from γδ-T or NK cells.. Nature communications. 2015 January ; 6 : 6025. doi:10.1038/ncomms7025.
Küçük C, Hu X, Jiang B, Klinkebiel D, Geng H, Gong Q, Bouska A, Iqbal J, Gaulard P, McKeithan TW, Chan WC. Global promoter methylation analysis reveals novel candidate tumor suppressor genes in natural killer cell lymphoma.. Clinical cancer research : an official journal of the American Association for Cancer Research. 2015 April ; 21 (7) : 1699-711. doi:10.1158/1078-0432.CCR-14-1216.
Bouska A, McKeithan TW, Deffenbacher KE, Lachel C, Wright GW, Iqbal J, Smith LM, Zhang W, Kucuk C, Rinaldi A, Bertoni F, Fitzgibbon J, Fu K, Weisenburger DD, Greiner TC, Dave BJ, Gascoyne RD, Rosenwald A, Ott G, Campo E, Rimsza LM, Delabie J, Jaffe ES, Braziel RM, Connors JM, Staudt LM, Chan WC. Genome-wide copy-number analyses reveal genomic abnormalities involved in transformation of follicular lymphoma.. Blood. 2014 March ; 123 (11) : 1681-90. doi:10.1182/blood-2013-05-500595.
Küçük C, Hu X, Iqbal J, Gaulard P, Klinkebiel D, Cornish A, Dave BJ, Chan WC. HACE1 is a tumor suppressor gene candidate in natural killer cell neoplasms.. The American journal of pathology. 2013 January ; 182 (1) : 49-55. doi:10.1016/j.ajpath.2012.09.012.
Perry AM, Warnke RA, Hu Q, Gaulard P, Copie-Bergman C, Alkan S, Wang HY, Cheng JX, Bacon CM, Delabie J, Ranheim E, Kucuk C, Hu X, Weisenburger DD, Jaffe ES, Chan WC. Indolent T-cell lymphoproliferative disease of the gastrointestinal tract.. Blood. 2013 November ; 122 (22) : 3599-606. doi:10.1182/blood-2013-07-512830.
Liu C, Iqbal J, Teruya-Feldstein J, Shen Y, Dabrowska MJ, Dybkaer K, Lim MS, Piva R, Barreca A, Pellegrino E, Spaccarotella E, Lachel CM, Kucuk C, Jiang CS, Hu X, Bhagavathi S, Greiner TC, Weisenburger DD, Aoun P, Perkins SL, McKeithan TW, Inghirami G, Chan WC. MicroRNA expression profiling identifies molecular signatures associated with anaplastic large cell lymphoma.. Blood. 2013 September ; 122 (12) : 2083-92. doi:10.1182/blood-2012-08-447375.
Cairns RA, Iqbal J, Lemonnier F, Kucuk C, de Leval L, Jais JP, Parrens M, Martin A, Xerri L, Brousset P, Chan LC, Chan WC, Gaulard P, Mak TW. IDH2 mutations are frequent in angioimmunoblastic T-cell lymphoma.. Blood. 2012 February ; 119 (8) : 1901-3. doi:10.1182/blood-2011-11-391748.
Küçük C, Iqbal J, Hu X, Gaulard P, De Leval L, Srivastava G, Au WY, McKeithan TW, Chan WC. PRDM1 is a tumor suppressor gene in natural killer cell malignancies.. Proceedings of the National Academy of Sciences of the United States of America. 2011 December ; 108 (50) : 20119-24. doi:10.1073/pnas.1115128108.
Iqbal J, Weisenburger DD, Chowdhury A, Tsai MY, Srivastava G, Greiner TC, Kucuk C, Deffenbacher K, Vose J, Smith L, Au WY, Nakamura S, Seto M, Delabie J, Berger F, Loong F, Ko YH, Sng I, Liu X, Loughran TP, Armitage J, Chan WC, International Peripheral T-cell Lymphoma Project.. Natural killer cell lymphoma shares strikingly similar molecular features with a group of non-hepatosplenic γδ T-cell lymphoma and is highly sensitive to a novel aurora kinase A inhibitor in vitro.. Leukemia. 2011 February ; 25 (2) : 348-58. doi:10.1038/leu.2010.255.
Iqbal J, Kucuk C, Deleeuw RJ, Srivastava G, Tam W, Geng H, Klinkebiel D, Christman JK, Patel K, Cao K, Shen L, Dybkaer K, Tsui IF, Ali H, Shimizu N, Au WY, Lam WL, Chan WC. Genomic analyses reveal global functional alterations that promote tumor growth and novel tumor suppressor genes in natural killer-cell malignancies.. Leukemia. 2009 June ; 23 (6) : 1139-51. doi:10.1038/leu.2009.3.
Total : 29
Selected Book Chapters
Genomik ve Uygulamaları (2023). Moleküler Biyoloji ve Genetik (Sağlık Alanında ve Biyoteknolojide Uygulamalar). TÜBA.
Lenfoma Genetiği (HL, NHL, B-cell lenfoma, KLL) (2019). HematoLog-Genetik. Galenos Yayınevi.
Total : 2
Projects
The Scientific and Technological Research Council of Turkey - TUBITAK - RD : REL Onkogeninin Foliküler Lenfoma Oluşumundaki Rolünün İncelenmesi, Finished
The Scientific and Technological Research Council of Turkey - TUBITAK - RD : Investigation of the Potential of the Mutations in Cell-free Circulating Tumor DNA as Biomarkers of Diagnosis and Chemo-immunotheraphy Resistance in Follicular Lymphoma Patients, Finished
The Scientific and Technological Research Council of Turkey - TUBITAK - RD : Identification of long non-coding RNAs that have potential to be novel diagnostic, prognostic biomarkers or therapeutic targets in mantle cell lymphoma through whole-transcriptome sequencing, Finished
Awards
- ASH Awards Supplement Program by American Society of Hematology, 2021
- 46th National Hematology Congress Abstract Award by Türk Hematoloji Derneği, 2020
- Trailblazers in Oncology Summit Manuscript Award by Türk Kanser Araştırma ve Savaş Kurumu Derneği, 2019
- Global Research Award by American Society of Hematology, 2018
- Young Investigator (GEBİP) Awards by Turkish Academy of Sciences (TÜBA), 2017
- Young Researcher Award (1st ranking) by XV. National Medical Biology and Genetics Congress, 2017
- Young Scientist (BAGEP) Award by Science Academy, 2015
- 2232 Yurda Dönüş Araştırmacı Dolaşım Programı by TUBITAK, 2014
- ASH Abstract Achievement Awards by American Society of Hematology, 2013
Academic Memberships
- TÜBA-Turkish Academy Of Science, 2017
- The Science Academy, Turkey, 2015
- Amerika Hematoloji Derneği (ASH), 2018
- Avrupa Hematoloji Derneği (EHA), 2017
- Lökosit Biyoloji Derneği (SLB), 2021
- Tıbbi Biyoloji ve Genetik Derneği (TBGDER), 2015
- Türk Hematoloji Derneği (THD), 2015
Contact

Kucuk Lab. on Lymphoid Cancer Genomics

