IBG-BIP Bioinformatics Platform
OVERVIEW
Welcome Who Are We ?
Izmir Biomedicine and Genome Center (IBG) is a pioneering research institution with the mission to expand the boundaries of life sciences and contribute to public health. One of the key components of the center is the Bioinformatics Platform (BIP), a unit specialized in the analysis of data generated from Next-Generation Sequencing (NGS) technologies, boasting extensive expertise in this field.
Our Mission
The primary mission of BIP is to strengthen and accelerate the scientific discovery process through NGS data analysis. To this end, we offer innovative and reliable bioinformatics solutions for researchers, as well as professionals operating in the biotechnology and pharmaceutical industries. Our platform employs cutting-edge algorithms and methodologies to analyze and interpret complex, multi-dimensional NGS datasets.
Our Services
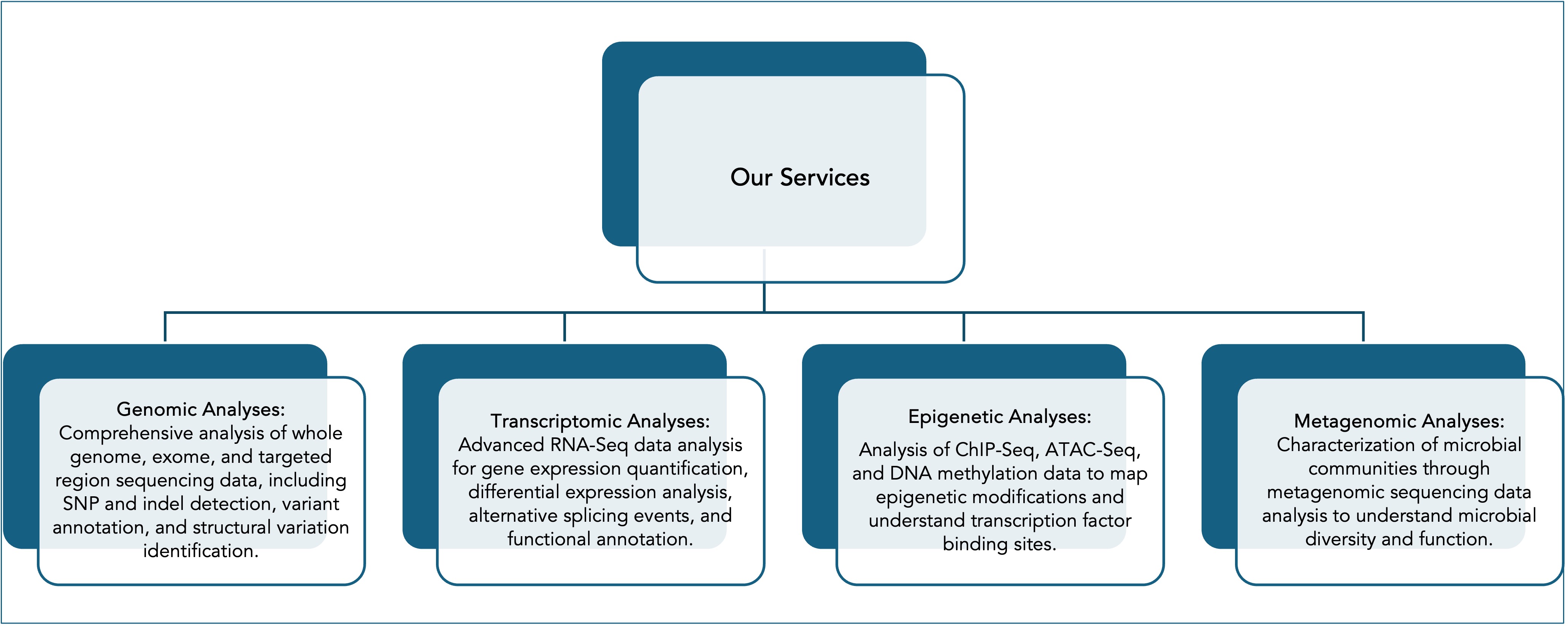
In addition to NGS data analysis, BIP provides a wide range of services, including customized consultancy. Detailed information about our services can be found on the services page of our platform. Some of the specialized services we offer include comprehensive analysis of data derived from RNA and DNA sequencing technologies, examination of transcriptome dynamics in various biological contexts, mapping of epigenetic modifications, and predicting the functional effects of variants.
Our Technical Infrastructure
Our platform is supported by a high-end technical infrastructure designed to handle large-scale NGS data analysis. This infrastructure includes high-performance computing systems and extensive data storage capacities. Thanks to this technical setup, our researchers meet high standards of accuracy and speed in bioinformatics data analysis.
Contact and Collaboration
Researchers and industry professionals interested in our bioinformatics analyses and consultancy services can contact us directly. We are here to support your scientific research and pave the way for new discoveries.
Personnel

IBG-BIP Bioinformatics Platform

Cihangir YANDIM
Visiting Researcher
cihangir.yandim@ibg.edu.tr

Necla KOÇHAN
Post-Doc Researcher
necla.kochan@ibg.edu.tr

Hüseyin GÜNER
Researcher
huseyin.guner@ibg.edu.tr
0 (232) 299 _ _
Former Personnel

Gökhan KARAKÜLAH
Platform Director
gokhan.karakulah@ibg.edu.tr
+90 232 299 41
00
(5051)
0 232 299 41 55

Tutku YARAŞ
Platform Director
tutku.yaras@ibg.edu.tr

İbrahim ŞENTÜRK
Research Group Member
ibrahim.senturk@ibg.edu.tr

Arif GÜRSOY
Research Group Member
arif.gursoy@ibg.edu.tr

Gökhan KARAKÜLAH
Research Group Member
gokhan.karakulah@ibg.edu.tr
+90 232 299 41
00
(5051)
0 232 299 41 55

Alirıza ARIBAŞ
R&D Personnel
aliriza.aribas@ibg.edu.tr

Uğur ÇABUK
MSc Student
ugur.cabuk@msfr.ibg.edu.tr

Necla KOÇHAN
Post-Doc Researcher
necla.kochan@ibg.edu.tr

Necati Kaan KUTLU
PhD Student
necatikaan.kutlu@ibg.edu.tr

Leman BİNOKAY
MSc Student
leman.binokay@ibg.edu.tr

Nurten Ezgi DOĞUDAN
Undergraduate Student
ezgi.dogudan@msfr.ibg.edu.tr

Fatih MAMAK
Undergraduate Student
fatih.mamak@ibg.edu.tr

Necati Kaan KUTLU
PhD Student
necatikaan.kutlu@ibg.edu.tr

Uğur DURA
Undergraduate Student
ugur.dura@ibg.edu.tr

Elif YILDIZ
MSc Student
elif.yildiz@ibg.edu.tr

Leman BİNOKAY
PhD Student
leman.binokay@ibg.edu.tr

Başak KESKİNOĞLU
Researcher
basak.keskinoglu@ibg.edu.tr

Fatih MAMAK
Undergraduate Student
fatih.mamak@ibg.edu.tr

Barış AKINCI
Researcher
baris.akinci@ibg.edu.tr
Contact

IBG-BIP Bioinformatics Platform